Abstract
New version of .NET Framework 4.6 with a lot of new features in it was release few weeks ago. One feature which got my attention is support for Simple Instruction Multiple Data (SIMD) vector instructions in the new 64-bit JIT compiler named RyuJIT. I wanted to see how easy is it to leverage this SIMD support and what performance improvements we can expect for code using it.
Introduction to SIMD
The SIMD types of instructions are as the name suggests instructions which take multiple operands and do some vector type operation with it (e.g. adding two vectors). They can work with different types (e.g. int, float, double, etc.) and the level of parallelism (vector size) depends on length of the register.
There are two types of vector instructions available in x86 compatible processors. The older one came with SSE2 instruction set which contains 128 bit registers for vector operations. The newer instructions are available in different type of AVX instruction sets. Depending on instruction set supported on processor (AVX, AVX2, AVX-512) it has register with size up to 512 bit. RyuJIT is trying to use the best one available for the CPU on which it is going to execute the code.
The support for SIMD was added just to the new RyuJIT compiler which at this moment works just for x64 programs. There were some claims that RyuJIT can be extended to other platforms as well but for now there will be no improvement for x86 programs. Second thing worth mentioning is that although the .NET team has target to run code using SIMD wrapper classes on the par with sequential code in case SIMD cannot be used, there are no there yet.
The way in which RyuJIT and .NET Framework adds support for SMID is by attaching JitIntristicAttribute to class or method. JIT then knows that it can ignore normal bytecode of the method and it can replace it with some special handling (e.g. SIMD instruction). There are few classes which are marked with JitIntristicAttribute so they should use SIMD optimisation (with proper JIT support):
- Vector2 – Fixed vector of two floating point numbers (e.g. to represent point in 2D space)
- Vector3 – Fixed vector of three floating point numbers (e.g. to represent point in 3D space)
- Vector4 – Fixed vector of four floating point numbers
- Vector<T> – Variable length vector of type T (128 bit – 256 bit depending on supported instruction set)
The first there fixed vector length types are part of BCL in .NET Framework 4.6 in System.Numerics namespace. The variable length vector type is available as NuGet package System.Numerics.Vectors version 4.1.0. For .NET Framework 4.5 both fixed and variable length type are in the same NuGet package of version 4.0.0.
Performance test
I created simple test which is multiplying two vectors in loop. The result amount of operations equals approximately to multiplying two matrices of size 100k x 100k. There are more complicated elaborate available but since I wanted to compare how the same code runs on different platforms and versions of .NET Framework I used just this simple test.
The test I created is trying to use all currently supported vector classes plus scalar implementation of vector multiplication as benchmark. I compiled the code in both .NET Framework 4.5.2 and in 4.6 and in both platforms x86 and x64. I ran the tests on laptop with Intel Core i7-2640M and on desktop with Intel Core i7-3770. Both of them have support for SSE-2 and AVX (which have 128 bit registers).
I don’t have access to any CPU which has support for AVX2 (256 bit) or AVX-512 (512 bit) unfortunately. If somebody has CPU which supports this instruction sets, let me know results of running the test application. I’d like to know the results for them as well. The source code for the application which I used is available on GitHub (taynes13/SimdTest).
The following table contains the test results. I did average of five test runs in order to avoid some random fluctuations. I normalised results for every CPU in order to be able to compare them, the value 100% is for x64 running on .NET Framework 4.6.
| .NET Version | Platform | Method | Avg. Time | Perf. Imp. | ||
|---|---|---|---|---|---|---|
| i7-2640M | i7-3770 | i7-2640M | i7-3770 | |||
| .NET 4.6 | x64 | Scalar | 00:09.3 | 00:08.2 | 100.00% | 100.00% |
| .NET 4.6 | x86 | Scalar | 00:09.4 | 00:08.2 | 98.90% | 100.16% |
| .NET 4.5.2 | x64 | Scalar | 00:09.5 | 00:08.3 | 97.35% | 99.44% |
| .NET 4.5.2 | x86 | Scalar | 00:09.3 | 00:08.2 | 100.00% | 100.35% |
| .NET 4.6 | x64 | VectorT | 00:05.6 | 00:03.5 | 167.02% | 237.01% |
| .NET 4.6 | x86 | VectorT | 01:11.7 | 01:02.6 | 12.94% | 13.16% |
| .NET 4.6 | x64 | Vector2 | 00:23.0 | 00:15.2 | 40.43% | 54.10% |
| .NET 4.6 | x86 | Vector2 | 00:21.5 | 00:17.7 | 43.18% | 46.56% |
| .NET 4.5.2 | x64 | Vector2 | 00:23.3 | 00:15.1 | 39.89% | 54.40% |
| .NET 4.5.2 | x86 | Vector2 | 00:21.2 | 00:17.7 | 43.79% | 46.52% |
| .NET 4.6 | x64 | Vector3 | 00:19.4 | 00:13.8 | 47.90% | 59.83% |
| .NET 4.6 | x86 | Vector3 | 00:20.0 | 00:16.9 | 46.31% | 48.72% |
| .NET 4.5.2 | x64 | Vector3 | 00:19.4 | 00:13.9 | 47.93% | 59.15% |
| .NET 4.5.2 | x86 | Vector3 | 00:20.4 | 00:17.8 | 45.59% | 46.27% |
| .NET 4.6 | x64 | Vector4 | 00:17.6 | 00:13.1 | 52.88% | 62.66% |
| .NET 4.6 | x86 | Vector4 | 00:25.9 | 00:23.3 | 35.88% | 35.34% |
| .NET 4.5.2 | x64 | Vector4 | 00:17.7 | 00:13.1 | 52.48% | 62.86% |
| .NET 4.5.2 | x86 | Vector4 | 00:26.0 | 00:23.5 | 35.74% | 35.00% |
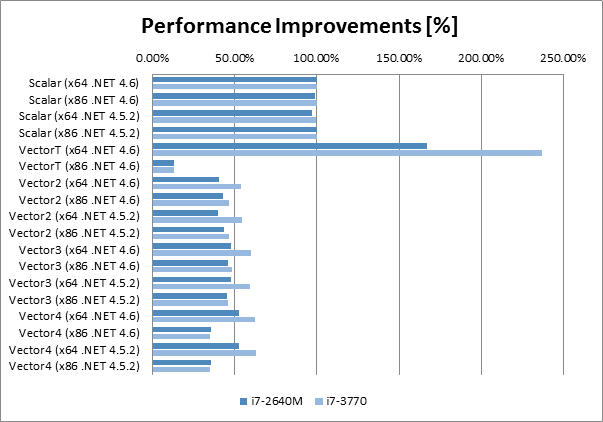
There are few interesting observations
- The Vector<T> running on the new RyuJIT (x64 .NET 4.6) with SIMD support has increased performance of multiplication by almost 2.5x on the desktop CPU (and bit more moderate improvement by 1.7x for laptop CPU).
- The same Vector<T> running on x86 .NET 4.6 was running 7.5x slower than scalar multiplication and almost 18x slower than the x64 RyuJIT using SIMD.
- All the other Vector2, Vector3 and Vector4 classes had very similar performance on both .NET 4.5.2 and .NET 4.6
- There was no performance improvement for x64 RyuJIT compiled code for Vector2, Vector3 and Vector4 classes. The actually performance degradation, for all the vector types the code runs on average 2x slower than scalar implementation
Summary
In my tests I can see that it is beneficial to use Vector<T> type if and only if we can ensure the application is going to run on x64 bit .NET Framework 4.6. If we are not sure whether the program is going to run on x86 or x64, it is better to avoid using Vector<T> because it will be by almost order of magnitude slower on x86 than scalar implementation. As I mentioned earlier the .NET team has target to run the programs using Vector classes without performance hit if run on x86 but we will have to wait for it.
Second thing which surprised me in my tests is that I haven’t seen performance improvements for any of the fixed length vectors (compared to x86 or event .NET Framework 4.5.2). This suggests that SIMD is disabled for the Vector classes in BCL of .NET Framework 4.6. I have seen everywhere on internet articles how RyuJIT adds support for SIMD but nothing about this support being switched off for fixed Vector classes. If somebody has some explanation, let me know, please.
To summarise my tests in one sentence, the support for RyuJIT is definitely promising, but one has to be careful where and how is the code going to be compiled and run (at least for now).